Research in the VanBuren LabDrought is the most pervasive issue we face in agriculture today, and water deficit is responsible for most of the global loss in yield and quality. Work in the VanBuren lab focuses on exploring the mechanisms that plants use to combat drought stress in natural and agricultural settings. We leverage large-scale, high-resolution datasets across space and time to model and predict the genetic elements, physiological signatures, and alleles controlling drought tolerance phenotypes. Our work centers on the C4 cereals maize, sorghum, teff, and various millets as well as numerous wild and model species with unique evolutionary innovations. In addition to our hypothesis-based research questions, we also specialize in developing computational approaches to sequence, analyze, and compare complex plant genomes.
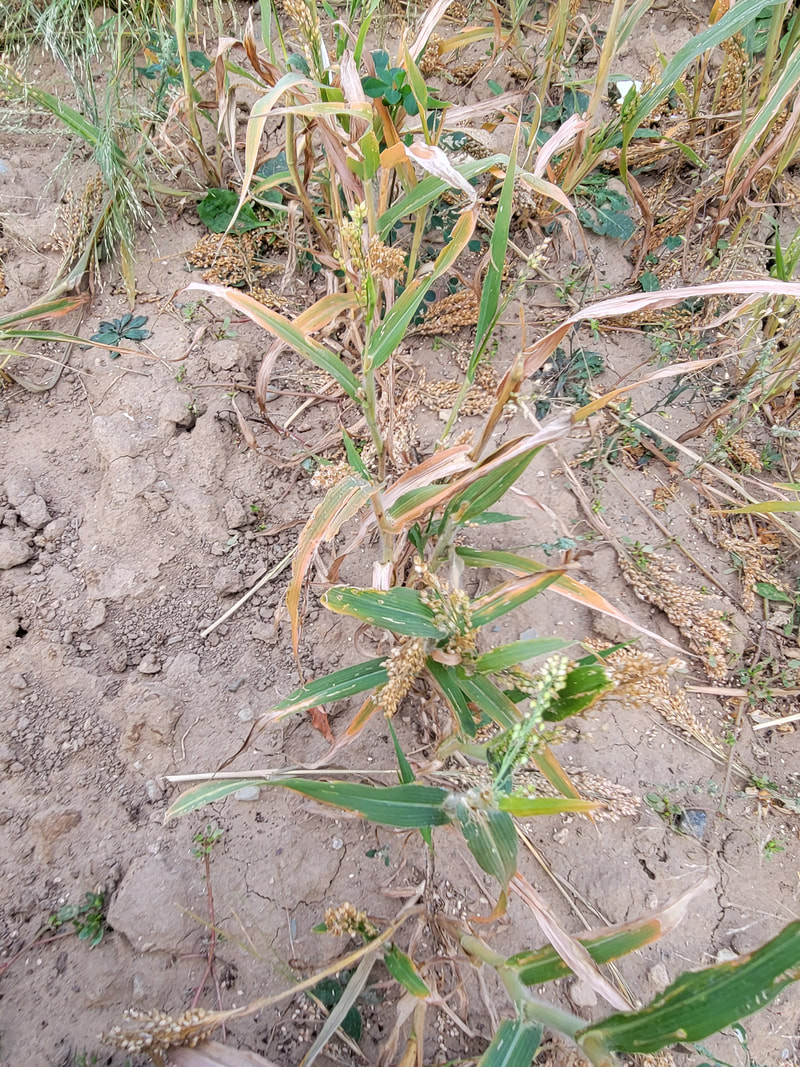
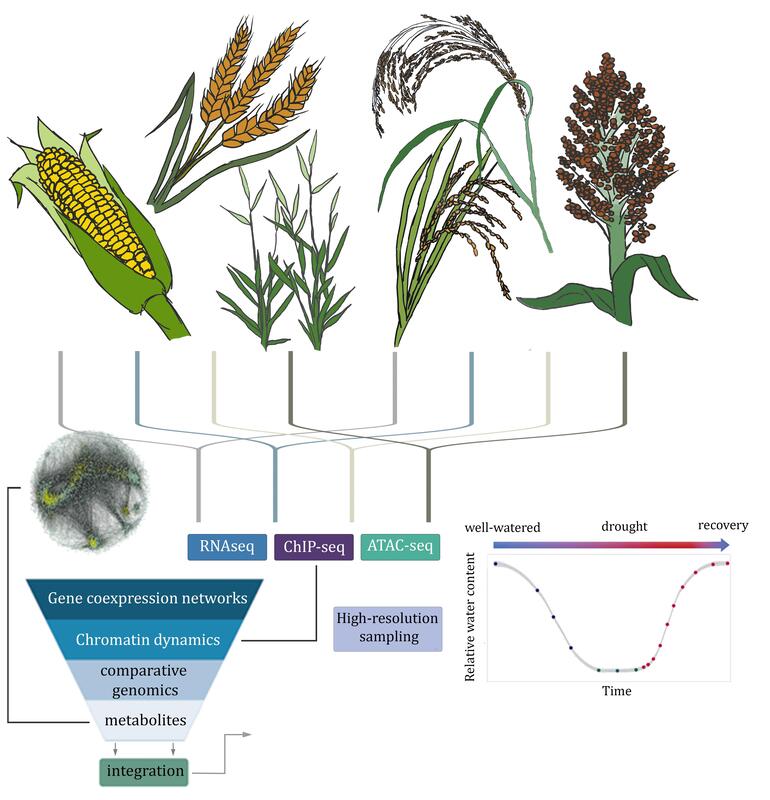
Improving the climate resilience of C4 cerealsGrasses provide the bulk of human nutrition and developing climate resilient cereals is central to maintaining global food security. Work in the lab focuses on understanding the genetic basis of stress tolerance using naturally resilient C4 cereals like teff, millets, and sorghum as well as the leading cereal maize. To identify the traits and genetic loci underlying stress tolerance in cereals, we use a combination of physiology, high-throughput phenotyping, quantitative genetics, expression dynamics, and predictive modeling-based approaches. We profile stress responses using highly controlled greenhouse, growth chamber, and field environments. The long-term goals of this research are to understand, model, and predict drought responses and to develop climate-resilient cereals.
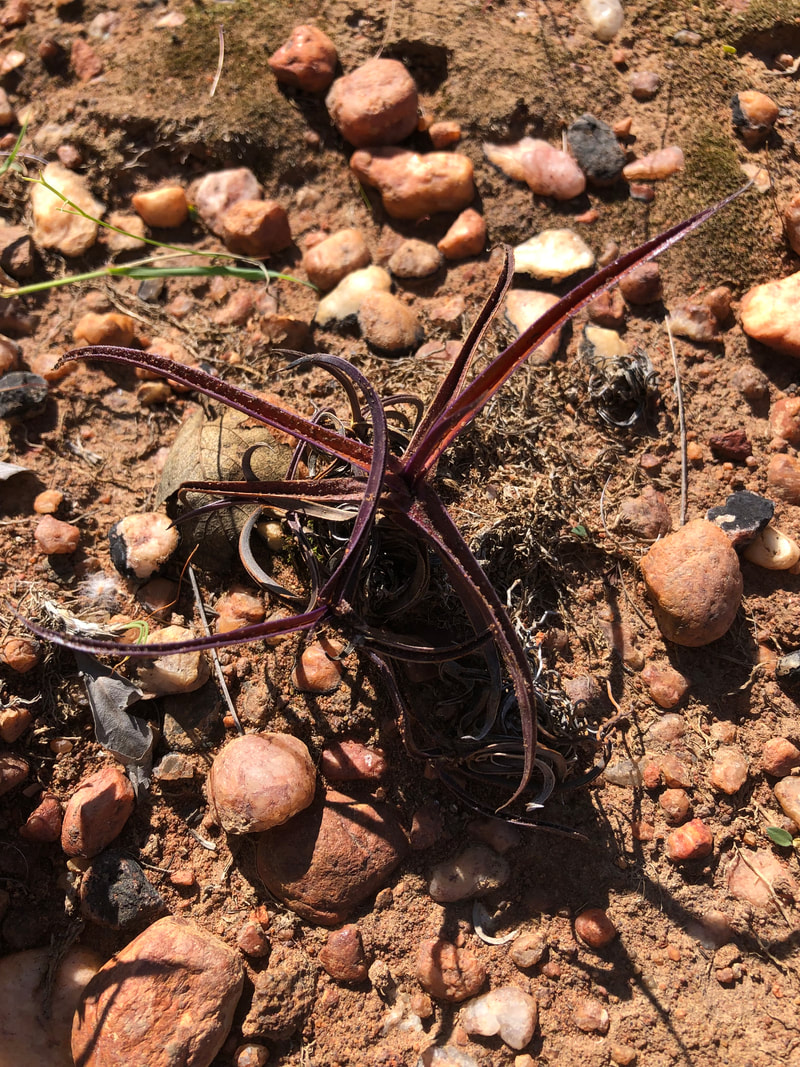
Understanding how life survives without waterAnhydrobioisis or life without water has evolved recurrently across the tree of life as a common adaptation to avoid drying without dying. A small group of plants can survive prolonged desiccation for months to years until the return of water. These so-called “resurrection plants” can desiccate to near complete anhydrobiosis (< 10% water content) for months to years with limited effects on cellular machinery, macromolecules, and plant viability. We aim to uncover the genetic basis of desiccation tolerance and identify components that may be used to improve plant resilience. To reach this goal, we utilize large-scale comparative genomic, transcriptomic, and physiologic datasets across diverse land plant and algae lineages spanning ~900 million years of divergence
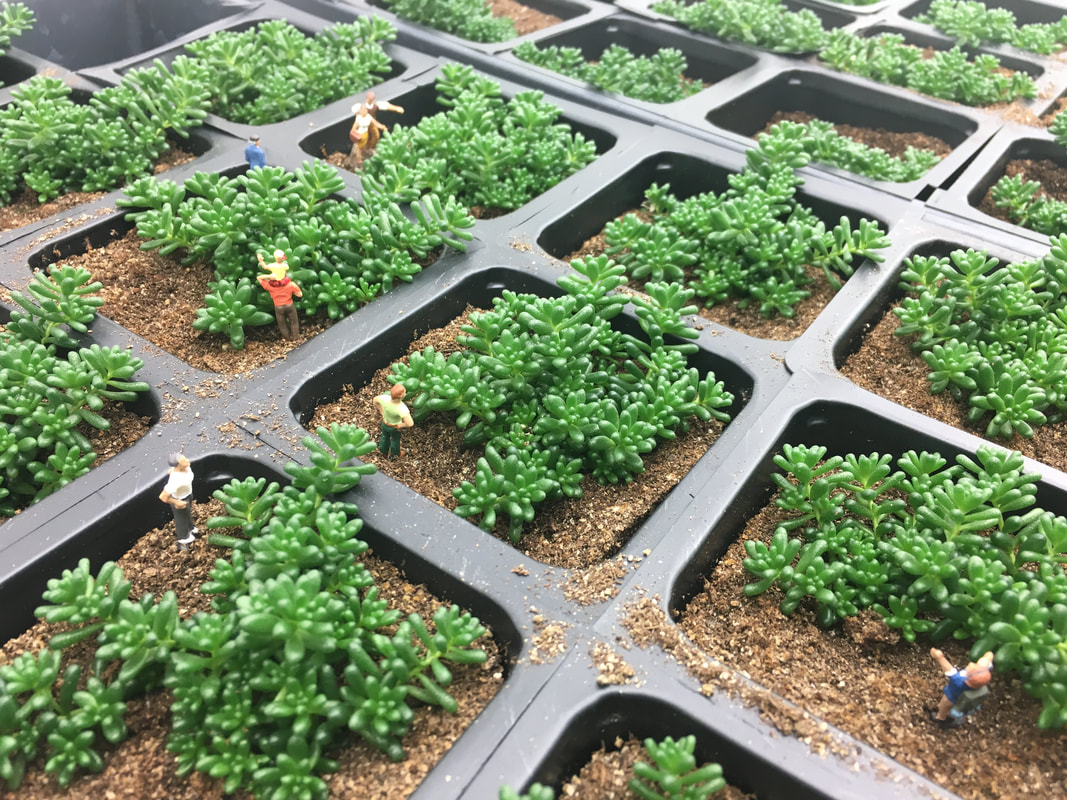
Engineering CAM photosynthesisCAM plants have evolved to survive in arid environments and use up to 80% less water than typical C3 species. Thus, engineering CAM into C3 or C4 crop plants could dramatically increase water use efficiency and drought tolerance, improving both yield and drought survivability. My lab uses a systems biology approach to explore the evolution of CAM photosynthesis in several distinct CAM lineages. Ongoing work focuses on the constitutive CAM plant pineapple and facultative CAM plants within the Sedum and Portulaca genera. We have generated high-resolution datasets across species, time, and development to better capture the dynamics of this complex pathway. The long-term goal of this project is to engineer the CAM pathway into C3 or C4 plants to improve water use efficiency
|